Benoît Serive
Entrepreneur & Marine pharmacognosy researcher
44 years old
Driving License
Nantes (44200) France
Professional Status
Freelancer
Just looking around
About Me
Laureate of the vocation award 2006
(Bleustein Blanchet Fundation)
Laureate 2013 - Marie Curie Outgoing Fellowship
OCEANCHArCoT programm
Member of the French Speaking Network of Metabolomics and Fluxomics
Member of the French Speaking Society of Pharmacognosy
Member of the American Society of Pharmacognosy
Member of the Association for the Sciences of Limnology and Oceanography
Reviewer for scientific peer-reviewed journals :
Marine Drugs, Algal Research
Invited Speaker at the 5th World Congress on Biotechnology (Valencia, Spain, June 2014)
In brief :
Voluntary, passionate and sensitive to human values, I want to use my skills to the search for molecules with high added value from marine biodiversity.
Golden prize of the most scholar, caring, and honest supervisor awarded to:
Prof. Ronald J. QUINN
(Eskitis Institute for Drug Discovery, Brisbane, Australia)
Mentor :
Emeritus Prof. Jean-Michel KORNPROBST
(University of Nantes, France)
(Bleustein Blanchet Fundation)
Laureate 2013 - Marie Curie Outgoing Fellowship
OCEANCHArCoT programm
Member of the French Speaking Network of Metabolomics and Fluxomics
Member of the French Speaking Society of Pharmacognosy
Member of the American Society of Pharmacognosy
Member of the Association for the Sciences of Limnology and Oceanography
Reviewer for scientific peer-reviewed journals :
Marine Drugs, Algal Research
Invited Speaker at the 5th World Congress on Biotechnology (Valencia, Spain, June 2014)
In brief :
Voluntary, passionate and sensitive to human values, I want to use my skills to the search for molecules with high added value from marine biodiversity.
Golden prize of the most scholar, caring, and honest supervisor awarded to:
Prof. Ronald J. QUINN
(Eskitis Institute for Drug Discovery, Brisbane, Australia)
Mentor :
Emeritus Prof. Jean-Michel KORNPROBST
(University of Nantes, France)
Poster displayed at the 8th International Conference on Marine Bioprospecting
Creation Date
07 mars 2017
Poster 9th European Conference on Marine Natural Products
Creation Date
31 août 2015
With the recent development of state-of-the-art technologies (e.g hyphenated MS techniques) and methodologies (e.gdereplication), the scientific community is interested in the exploration of poorly chemically studied bioresources. The high diversity of interacting phytoplankton species suggests an important and highly diverse chemical repertoire (e.gisoprenoids, toxins, polysaccharides, PUFAs, oxylipins, phycobiliproteins) which may inspire applications in health, nutrition and biotechnology. Biosynthesis of these metabolites is strongly dependent upon their environment/culture conditions which may be investigated using OMICS approaches. In microalgae, a major bottleneck isthe difficulty in extracting deeply inaccessible molecules, an important issue that demands adapted solutions prior to considering High-Throughput Screening (HTS). Bioactive minority metabolites may pass unnoticed on spectra and thus require special attention. The extraction of metabolites may prove difficult due to the presence of highly resistant cell walls (Phaeodactylumtricornutum), or of exopolysaccharidic secretions surrounding the cell membrane (Porphyridiumpurpureum). The Mix Mill process (vibrating microbeads) which gave excellent extraction yields without chemical alteration of the analytes) and is fully compatible with HPLC and LC-MS analysis was optimised. Being accurate, simple to operate, rapid, safe and preserving sensitive molecules, makes the Mix Mill process suitable for the screening of microalgalchemodiversity. This methodology was applied in the Photomer, and currently in OCEANOMICs and OCEANCHArCoT programs, all being dedicated to the identification of new marine metabolites with high added value. Finally, this methodology represents a significant improvement in the field of OMICS studies from microalgae, as it provides the most representative estimate of their exploitable chemical diversity.
Annual general meeting of BIOCHIMAR group
Creation Date
08 nov. 2013
Development and optimization of a metabolite extraction process for the high throughput screening of microalgal chimiodiversity
Algae : an incredible reservoir of applications as their biodiversity
Creation Date
01 mars 2012
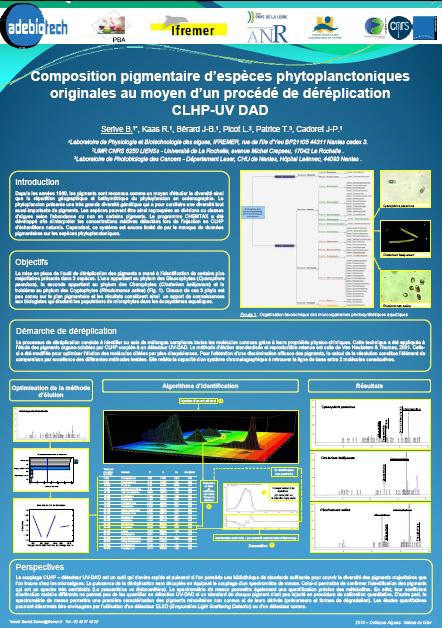
Study of microalgal pigments in oceanography and biotechnology through a process of dereplication HPLC-UV DAD
Creation Date
20 oct. 2011
Collection of marine tropical organisms in scuba-diving - From the study of biodiversity to the emergence of new drugs
Creation Date
18 mars 2011
A pigment composition of original phytoplankton species by means of a method of HPLC-UV DAD dereplication